Rooting and bootstrap
Taxonomy
Serratia prometaculans:cellular organisms, Bacteria, Proteobacteria,Gammaproteobacteria,Enterobacterales,Yersiniaceae.
Yersinia pestis: cellular organisms, Bacteria, Proteobacteria, Gammaproteobacteria, Enterobacterales, Yersiniaceae, Yersinia pseudotuberculosis complex
Escherichia coli: cellular organisms, Bacteria, Proteobacteria, Gammaproteobacteria, Enterobacterales, Enterobacteriaceae
Proteus mirabilis: cellular organisms, Bacteria, Proteobacteria, Gammaproteobacteria, Enterobacterales, Morganellaceae
Haemophilus influenzae: cellular organisms, Bacteria, Proteobacteria, Gammaproteobacteria, Pasteurellales, Pasteurellaceae
Pasteurella multocida: cellular organisms, Bacteria, Proteobacteria, Gammaproteobacteria, Pasteurellales, Pasteurellaceae
Shewanella denitrificans: cellular organisms, Bacteria, Proteobacteria, Gammaproteobacteria, Alteromonadales, Shewanellaceae
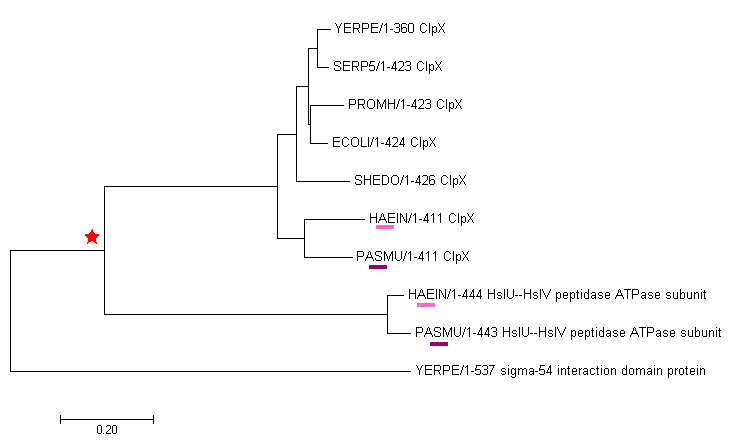
Here we can see phylogenetic tree of selected bacteria, builded with Muscle with defaults and NJ method. All of the proteins selected with sufficient e-values.
2 types of proteins are divided: ClpX protein and HsIU--HsIV peptidase ATPase subunit. It may be a result of the duplication in the position marked with star. So HsIU--HsIV peptidase ATPase
subunit from Pasteurella multocida and HsIU--HsIV peptidase ATPase subunit from Haemophilus influenzae are paralogs with ClpX proteins in that organisms. Also, we can see the orthologs
(Yersinia pestis and Serratia proteamaculans ClpX, Shewanella denitrificans and Escherichia coli ClpX, Haemophilus influenzae and Pasteurella multocida ClpX.
All of them appeared as a result of speciation). Also, we can see that another duplication happens, when sigma-54 interaction domain protein in emerged Yersinia pestis.
© Gumerov Ruslan, 2017
