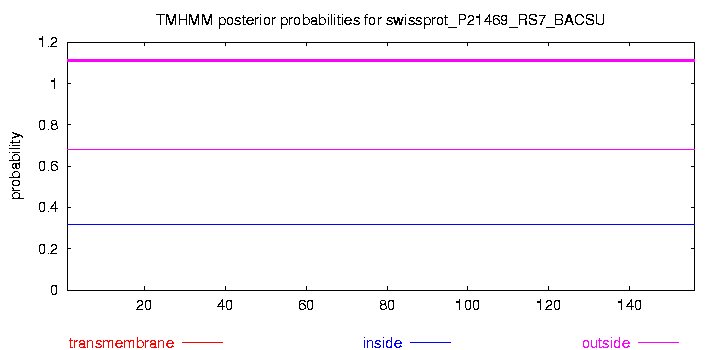
# swissprot_P21469_RS7_BACSU Length: 156 # swissprot_P21469_RS7_BACSU Number of predicted TMHs: 0 # swissprot_P21469_RS7_BACSU Exp number of AAs in TMHs: 0 # swissprot_P21469_RS7_BACSU Exp number, first 60 AAs: 0 # swissprot_P21469_RS7_BACSU Total prob of N-in: 0.31741 swissprot_P21469_RS7_BACSU TMHMM2.0 outside 1 156

# plot in postscript,
script for making the plot in gnuplot,
data for plot
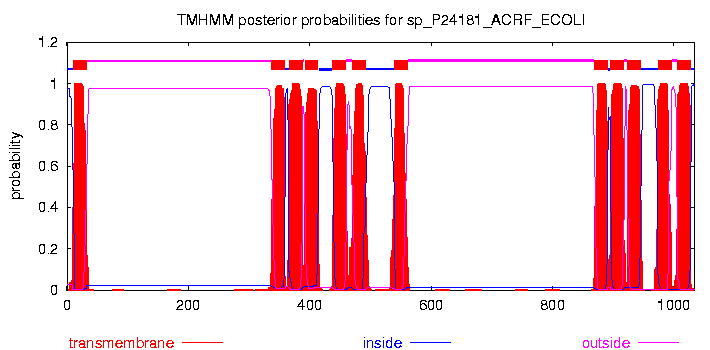
# sp_P24181_ACRF_ECOLI Length: 1034 # sp_P24181_ACRF_ECOLI Number of predicted TMHs: 12 # sp_P24181_ACRF_ECOLI Exp number of AAs in TMHs: 260.54891 # sp_P24181_ACRF_ECOLI Exp number, first 60 AAs: 21.70611 # sp_P24181_ACRF_ECOLI Total prob of N-in: 0.97501 # sp_P24181_ACRF_ECOLI POSSIBLE N-term signal sequence sp_P24181_ACRF_ECOLI TMHMM2.0 inside 1 10 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 11 33 sp_P24181_ACRF_ECOLI TMHMM2.0 outside 34 336 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 337 359 sp_P24181_ACRF_ECOLI TMHMM2.0 inside 360 365 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 366 388 sp_P24181_ACRF_ECOLI TMHMM2.0 outside 389 391 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 392 414 sp_P24181_ACRF_ECOLI TMHMM2.0 inside 415 437 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 438 460 sp_P24181_ACRF_ECOLI TMHMM2.0 outside 461 469 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 470 492 sp_P24181_ACRF_ECOLI TMHMM2.0 inside 493 539 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 540 562 sp_P24181_ACRF_ECOLI TMHMM2.0 outside 563 868 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 869 891 sp_P24181_ACRF_ECOLI TMHMM2.0 inside 892 895 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 896 918 sp_P24181_ACRF_ECOLI TMHMM2.0 outside 919 922 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 923 945 sp_P24181_ACRF_ECOLI TMHMM2.0 inside 946 973 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 974 996 sp_P24181_ACRF_ECOLI TMHMM2.0 outside 997 1005 sp_P24181_ACRF_ECOLI TMHMM2.0 TMhelix 1006 1028 sp_P24181_ACRF_ECOLI TMHMM2.0 inside 1029 1034
# plot in postscript,
script for making the plot in gnuplot,
data for plot