This minireview details a bioinformatic study of the Austwickia chelonae genome (GCF_003391095.1). Analyses included protein length and GC-content distributions, quantification of genomic features across its chromosome and plasmid, and the proportional length these features occupy. The results depict a classic bacterial architecture: a core chromosome encoding essential machinery and a small, accessory plasmid. Notably, a diphtheria toxin-like gene was identified, marking its first discovery outside the genus Corynebacterium and suggesting a potential virulence mechanism acquired via horizontal gene transfer.
Austwickia chelonae is a filamentous, Gram-positive bacterium first isolated in 1995 from a snapping turtle (Elseya sp.) with skin lesions at Perth Zoo, Australia [2]. The bacterium is primarily known for causing dermatological diseases, such as cutaneous granulomas, in reptiles. Notably, it infects endangered species, including the crocodile lizards (Shinisaurus crocodilurus Ahl, 1930) from Guangdong Luokeng, China [1]. Genomic analysis has revealed that A. chelonae possesses a diphtheria-like toxin gene, which encodes one of the most potent biotoxins known to halt protein synthesis [2]. This finding, reported in 2018, marked the first identification of a diphtheria-like toxin outside the genus Corynebacterium [3]. Antimicrobial susceptibility testing indicates that A. chelonae is sensitive to cephalothin, minocycline and ampicillin, but resistant to kanamycin, gentamicin, streptomycin and clarithromycin, suggesting potential treatment avenues for infected reptiles [1].
| Domain | Bacteria |
|---|---|
| Kingdom | Bacillati |
| Phylum | Actinomycetota |
| Class | Actinomycetes |
| Order | Micrococcales |
| Family | Dermatophilaceae |
| Genus | Austwickia |
| Species | Austwickia chelonae |
The Austwickia chelonae genome (assembly GCF_003391095.1) was obtained from the National Center for Biotechnology Information (NCBI) Genome Assembly database [S1]. The following files from this assembly were downloaded and used:
All data were imported into Google Sheets for analysis.
The length of each protein was calculated in amino acids (aa). From this dataset, the maximum and minimum protein lengths in the genome were identified. To visualise the distribution, a histogram was constructed with a bin width of 20 aa. The x-axis represents protein length (aa), and the y-axis represents the frequency (absolute count) of proteins within each bin.
The GC content was calculated for each protein-coding sequence (CDS) and expressed as a percentage. The maximum and minimum GC content values in the genome were identified. To visualise the distribution, a histogram was constructed with a bin width of 1%. The x-axis represents GC content (%), and the y-axis represents the frequency (absolute count) of CDS within each bin.
The absolute counts of each gene type — including protein-coding genes, pseudogenes, and RNA genes (tRNA, tmRNA, rRNA, ncRNA) — were tallied for each of the two replicons (chromosome and plasmid) in the Austwickia chelonae genome.
Each genomic feature category — including protein-coding genes, pseudogenes, RNA genes (tRNA, tmRNA, rRNA, ncRNA), and intergenic regions — was tallied for each of the 2 replicons (chromosome and plasmid) in the Austwickia chelonae genome. These counts were then expressed as percentages of the total sequence length of their respective replicons.
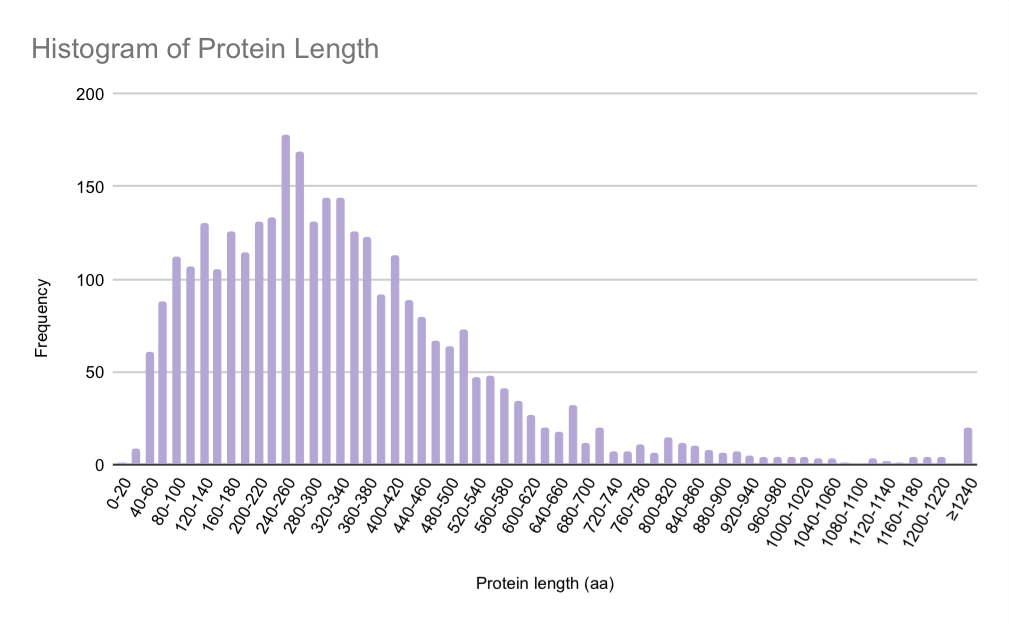
Figure 1 presents the distribution of protein lengths in the Austwickia chelonae genome. The histogram is right-skewed, with a pronounced peak at approximately 180 proteins in the 240-260 amino acid range. Most proteins are shorter than 700 amino acids. A long tail of the distribution extends to lengths exceeding 1,240 amino acids, where the frequency falls significantly to around 20 proteins per bin.

These results align with the typical average length of bacterial proteins, which is around 320 amino acids [4]. This suggests that the proteome of A. chelonae does not rely on a genome-wide trend toward exceptionally large proteins.
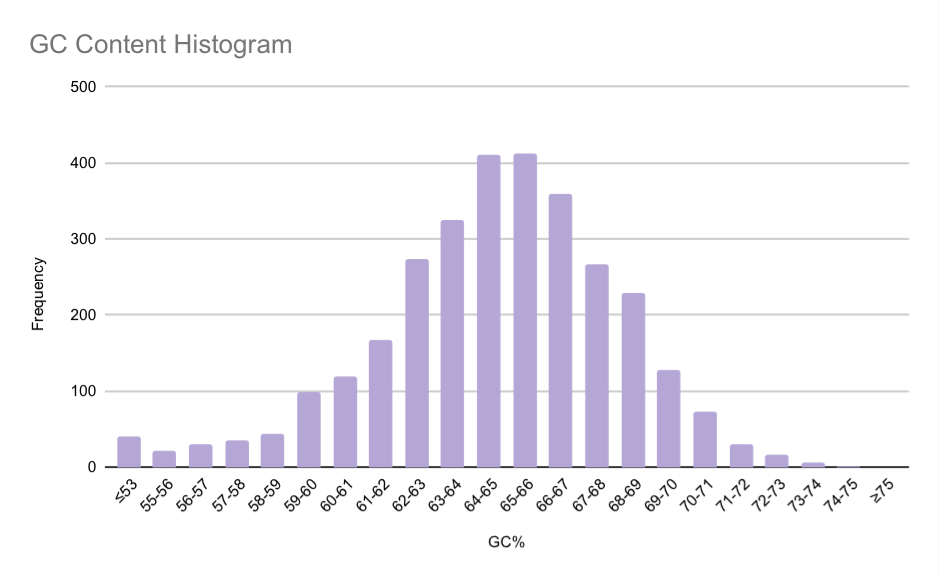
The GC content distribution of Austwickia chelonae CDS is shown in Figure 2. The histogram reveals a slightly left-skewed distribution, indicating that while most genes have a high GC content, a tail of genes with lower GC content still exists. The peak value is in the 65-66% bin, which contains 413 genes, representing the most common nucleotide composition for the coding genome. The overall range of GC content is wide, extending from 39% to 75%. Notably, only 41 genes were found to have a GC content below 51%. To improve the visualisation of the main distribution, gene counts for GC values ≤53% were consolidated into a single bin.
A high GC content is associated with greater DNA thermostability, as guanine-cytosine base pairs form 3 hydrogen bonds instead of the 2 in adenine-thymine pairs [5]. The elevated GC content observed in A. chelonae may therefore contribute to genomic stability. This could be advantageous in its ecological niche, potentially involving exposure to the variable body temperatures of its reptilian hosts.
The further analyses presented in this section include an examination of the distribution of RNA gene types. Table 2 lists the RNA species identified in the A. chelonae genome and describes their primary functions.
| RNA type | Primary Function |
|---|---|
| Transfer RNA (tRNA) | Transports amino acid to the growing polypeptide chain during translation, as specified by the mRNA codon [6]. |
| Transfer-messenger RNA (tmRNA) | Key component for bacterial translation quality control system. It rescues ribosomes stalled on damaged mRNA molecules, facilitating the release of the ribosome and the degradation of the incomplete protein [7]. |
| Ribosomal RNA (rRNA) | Catalytic and structural core of the ribosome [8]. |
| Non-coding RNA (ncRNA) | Regulates gene expression [9]. |
The distribution of different gene types in the A. chelonae genome between its chromosome (NZ_CP031447.1) and plasmid (NZ_CP031448.1) is shown in Table 3. The vast majority of genes are located on the chromosome, which harbors 3,214 features, accounting for 99.88% of the total gene count. These include 3,110 protein-coding genes, 50 pseudogenes, and a full complement of RNA genes (45 tRNA, 1 tmRNA, 6 rRNA, and 2 ncRNA). In stark contrast, the plasmid carries only 4 protein-coding genes and lacks other features, such as pseudogenes or RNA genes. This results in a total of 3,218 genes for the organism. The minimal contribution of the plasmid (0.12%) supports its classification as an auxiliary genetic element in the organism.
| Gene type | Chromosome (NZ_CP031447.1) | Plasmid (NZ_CP031448.1) | Total |
|---|---|---|---|
| Proteins | 3110 | 4 | 3114 |
| Pseudogenes | 50 | 0 | 50 |
| tRNA | 45 | 0 | 45 |
| tmRNA | 1 | 0 | 1 |
| rRNA | 6 | 0 | 6 |
| ncRNA | 2 | 0 | 2 |
| Total | 3214 | 4 | 3218 |
Table 4 presents a quantitative overview of genomic features across the 2 replicons of the Austwickia chelonae genome: chromosome (NZ_CP031447.1) and plasmid (NZ_CP031448.1). As is typical for bacteria, the chromosome contains the complete set of essential genetic elements. This includes all RNA gene types — transfer (tRNA), transfer-messenger RNA (tmRNA), ribosomal RNA (rRNA), and non-coding RNA (ncRNA) — along with a small proportion of pseudogenes (0.83% of all chromosomal features) and a substantial fraction of intergenic regions (11.34% of all chromosomal features). In contrast, the plasmid lacks these core RNA elements and contains no pseudogenes. It consists solely of protein-coding sequences (81.48% of plasmid length) and a significant proportion of intergenic space (18.52% of plasmid length). This distribution confirms the plasmid's role as an accessory element, potentially carrying adaptive genes, while the chromosome encodes the essential systems required for cellular viability.
| Genomic Feature Category | Chromosome (NZ_CP031447.1) | Plasmid (NZ_CP031448.1) |
|---|---|---|
| Proteins | 87.46% | 81.48% |
| Pseudogenes | 0.83% | 0.00% |
| tRNA | 0.10% | 0.00% |
| tmRNA | 0.01% | 0.00% |
| rRNA | 0.26% | 0.00% |
| ncRNA | 0.01% | 0.00% |
| Intergenic regions | 11.34% | 18.52% |
| Length (bp) | 3627619 | 2467 |
Austwickia chelonae encodes a diphtheria-like toxin (WP_162873017.1; length: 603 aa [S5]), a potent bacterial toxin previously characterised only within the genus Corynebacterium [2, 3]. This finding represents the first report of this toxin family outside that genus, raising significant questions about its origin and function. Future research should be focused on a detailed analysis of the toxin’s genomic context, including the prediction of its potential operon structure. This analysis would provide crucial insights into its genomic regulation and potential acquisition via horizontal gene transfer.
S1. NCBI Genome Assembly
The genome assembly of Austwickia chelonae (GCF_003391095.1) used as the
source for all genomic sequences (CDS), and genomic features table.
https://ftp.ncbi.nlm.nih.gov/genomes/all/GCF/003/391/095/GCF_003391095.1_ASM3
39109v1/
S2. NCBI Taxonomy Database
Source for the complete taxonomic lineage of Austwickia chelonae
(NCBI:txid100225).
https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?command=show&mod
e=node&id=100225&lvl=
S3. Coding sequences (CDS) table
All annotated coding sequences (CDS) from the Austwickia chelonae genome, used
for analyses.
CDS from genome of Austwickia chelonae
S4. Genomic feature table
A table containing the genomic feature table for Austwickia chelonae and the
following derived analytical sheets:
S4.1: CDS. Full genomic feature table.
S4.2: Gene. Genes features (filtered CDS table).
S4.3: CDS_with_protein. CDS features with proteins (filtered CDS table).
S4.4: CDS_without_protein. CDS features with pseudogenes (filtered CDS
table).
S4.5: Prot_length_hist. Histogram and statistics for protein length.
S4.6: GC_hist. Histogram and statistics for GC content.
S4.7: Per_replicons. Distribution of genomic features across replicons.
S4.8: Percents_of_length. Percentage of replicon length occupied by each
feature type.
Genomic feature of Austwickia chelonae
S5. NCBI Protein Database
Source for the diphtheria toxin-like protein identified in Austwickia chelonae
(Accession: WP_162873017.1).
https://www.ncbi.nlm.nih.gov/protein/WP_162873017.1
I thank my family for their unequalled financial and moral support throughout my education. A huge thanks also goes to my friend Yaroslava for all her help figuring out the genomic feature tables.